Diverse capsid maturation prohead protease mutations enable
Download scientific diagram | Diverse capsid maturation prohead protease mutations enable evasion of Type I CBASS immunity. (A) Cartoon schematic representation of parallel forward genetic screening methods for CBASS-resistant mutant phage selection in solid (method #1) and liquid media (method #2). (B) Predicted structure of the Drexlerviridae phage prohead protease with CBASS escaper phage mutations identified in this study depicted in red spheres. Numbers represent the number of phage mutants isolated with each mutation. (C) Sequencing analysis of Drexlerviridae phage prohead proteases highlighting substitution mutations by vertical boxes and the phage(s) which harbor each mutation denoted by horizontal boxes. Drexlerviridae subfamily groups are labeled on the Y-axis and cartoon representations below depict predicted protein secondary structure in those regions. (D) Solid media plaque assays showing CBASS escaper phage isolate resistance phenotypes in cells expressing the WT CBASS operon which they were selected against or catalytically inactive CD-NTase CBASS operons which contain two Asp to Ala (DDAA) substitutions in the nucleotidyltransferase active site. from publication: Convergent mutations in phage virion assembly proteins enable evasion of Type I CBASS immunity | CBASS is an anti-phage defense system that protects bacteria from phage infection and is evolutionarily related to human cGAS-STING immunity. cGAS-STING signaling is initiated by viral DNA but the stage of phage replication which activates bacterial CBASS remains unclear. | Phage, Bacteriophage and Operon | ResearchGate, the professional network for scientists.

Biophysics of viral infectivity: matching genome length with
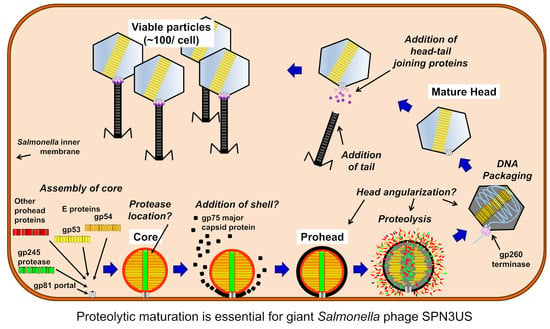
Viruses, Free Full-Text

Structure and function of bacteriophage T4

The Prohead-I structure of bacteriophage HK97: implications for
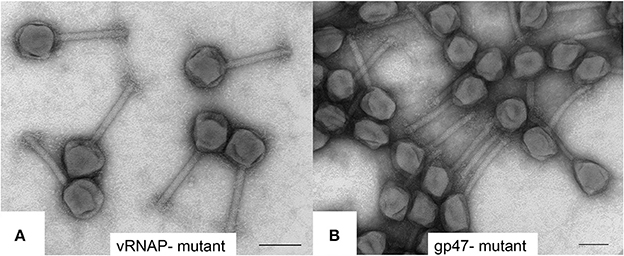
Frontiers To Be or Not To Be T4: Evidence of a Complex

Functional Domains of the HK97 Capsid Maturation Protease and the

Amy S.Y. Lee's research works

Structure and function of bacteriophage T4
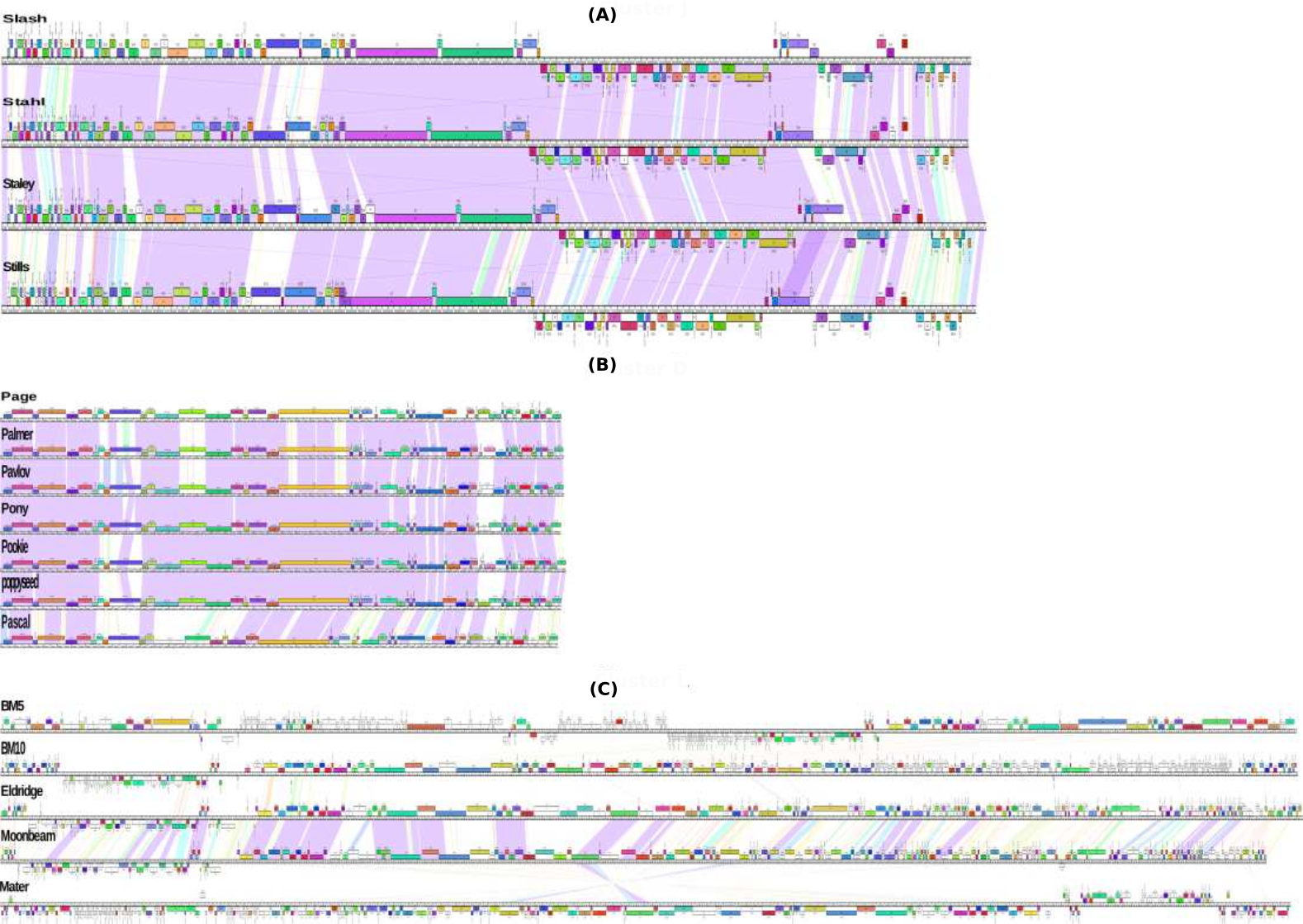

Characterization and comparative genomic analysis of virulent and

Samuel J. Hobbs's research works Dana-Farber Cancer Institute, Boston (DFCI) and other places

Desmond Richmond-Buccola's research works Harvard Medical School, MA (HMS) and other places

Samuel J. Hobbs's research works Dana-Farber Cancer Institute, Boston (DFCI) and other places









